Clinical Laboratory
The Steve and Cindy Rasmussen Institute for Genomic Medicine Clinical Laboratory at Nationwide Children’s Hospital performs high complexity molecular genetic analysis, cytogenetic analysis and advanced genomic testing.
The Steve and Cindy Rasmussen Institute for Genomic Medicine Clinical Laboratory also serves as the testing reference laboratory for the Acute Lymphoblastic Leukemia and Neuroblastoma Children's Oncology Groups, performing molecular and cytogenetic analyses for Children’s Oncology Group (COG) clinical trial patients.
Accreditation
The laboratory is accredited by the College of American Pathologists (CAP; #8162772) and certified by the Clinical Laboratory Improvement Amendments (CLIA; #36D2131848).
Faculty and Staff
The laboratory is led by eight clinical directors and one technical director. Its staff is composed of more than 30 technologists and technicians, 10 order entry and quality assurance staff, a lab operations manager, six lab supervisors, four clinical laboratory genetic counselors, five clinical bioinformatics personnel, six variant scientists and analysts, three clinical laboratory fellows and four clerical support staff.
The lab’s accessioning and quality assurance teams ensure that all clinical orders are recorded appropriately in its laboratory information management system (LIMS) and clinical reporting system. Accessioning personnel are the first people to greet you over the phone and are the receiving staff for incoming samples and testing requests. The accessioning team generates barcodes for the samples our lab receives and coordinates with the processing and testing lab sections to ensure testing is performed in a timely manner. The quality team verifies testing orders and documentation for accuracy as well as monitoring turnaround times, equipment documentation, training and competency, and procedure updates. They ensure we are following Clinical Laboratory Improvement Amendments (CLIA) guidelines and checklist requirements provided by the College of American Pathologists (CAP) as well as safety guidelines from regulatory bodies such as the United States Occupational Safety and Health Administration (OSHA).
Resources and Services
Our lab is fully equipped with instrumentation and computational resources to perform a variety of molecular, cytogenetic and advanced genomic analyses. It includes facilities for sample accessioning, tissue culture, cytogenetic analysis, fluorescence in situ hybridization (FISH), nucleic acid extraction, nucleic acid amplification, microarray analysis, methylation analysis, single gene sequencing, gene fusion analysis, next-generation sequencing and local freezer storage of patient samples.
Click here to view a list of all clinically offered lab tests for external clients.
A nucleic acid extraction is the act of releasing DNA/RNA from cells in a sample for use in tests and analytical procedures. Extracting the maximum amount of high-quality DNA and RNA from each sample is a critical step in the overall workflow for molecular diagnostics. High quality extractions reduce the risk that a test will fail to produce an interpretable result.
 Common Sample Types
Common Sample Types
- Blood
- Bone marrow
- Tissue (Paraffin Embedded and Snap Frozen)
- Cheek (buccal) swabs
- Saliva
Why Is DNA/RNA Isolated?
- DNA is the blueprint, recipe or code used to make a person. These directions determine everything that make a person unique, including hair color, eye color and height.
- RNA translates, or decodes, the directions given by DNA so the body and its cells can read and use them.
- If there’s a typo in these directions — a missing “word” or an added “word” — the wrong information can be sent to a person’s body.
- DNA copying mistakes and “mistyped” directions can cause complications with a person's development called mutations.
- DNA and RNA are isolated during diagnostic tests and analyses to get a view of what parts of the DNA blueprint are incorrect and causing harm to the person.

How are Nucleic Acids Isolated?
- Releasing the DNA and RNA from a cell requires breaking the cell open. The process of breaking the cell open is called cell disruption or cell lysis. This can be done through cell lysis buffer, disruption or sonicating.
- Solid‐phase extractions, or spin column extractions, are commonly used for DNA and RNA isolations, concentrating the sample for analysis. In solid-phase extractions, whole blood, bone marrow or white blood cells are applied to a column in a low pH, high salt and detergent solution. The nucleic acid, in solution, adsorbs to the solid matrix and is washed. Using a low salt solution, nucleic acid is then released from the matrix.
- Magnetic bead purification methods are also used for nucleic acid isolations. Whole blood, bone marrow, saliva, or white blood cells are lysed and bound to magnetic beads. Magnetic beads are positively charged at a low pH, attracting and binding the negatively charged nucleic acid. Nucleic acid is washed and subsequently removed from the magnetic beads, leaving behind a pure product.
Tools for Evaluating DNA/RNA Extractions

Fluorometer – Uses fluorescent binding dyes that bind to DNA or RNA. Bound dyes absorb specific wavelengths of light and emit light of another wavelength depending on the concentration of the sample.
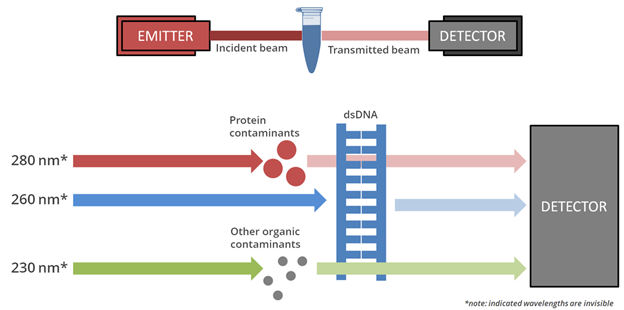
Spectrometer – Uses certain wavelengths of light to measure how much light is absorbed by the specimen.
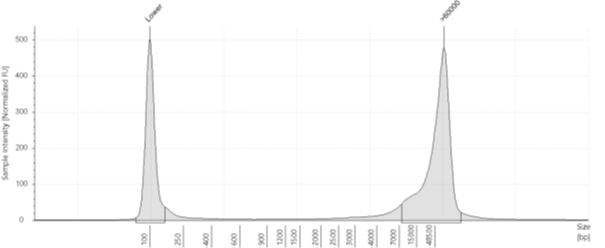
Capillary Analysis – Uses capillary electrophoresis and fluorescent detection to assigns numerical integrity values using a voltage gradient.
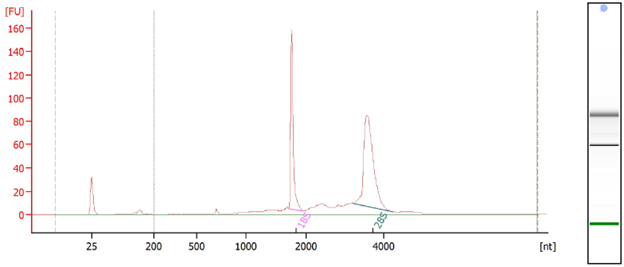
Gel Electrophoresis – Compares genomic weight to the ladder (a standard reference that contains nucleic acid fragments of known lengths).
Inherited Diseases
Our clinical laboratory offers numerous DNA based tests that can be used to diagnose genetic disorders and identify patients that are at risk of carrying genes associated with certain diseases. We offer a wide array of molecular tests, including sanger sequencing, fragment analysis, methylation analysis and targeted mutation analysis. Our genetic tests for inherited disease include:
- Carrier and diagnostic testing for cystic fibrosis and fragile X
- Follow up testing from newborn screening for cystic fibrosis, Krabbe disease and Hurler syndrome
- Methylation testing for Prader Willi and Angelman syndromes
- Targeted testing for known disease-associated familial variants
- Bone marrow engraftment monitoring
Cancer
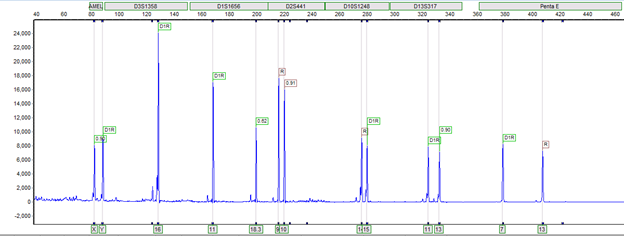
Our laboratory also offers targeted oncology DNA and RNA based testing. This includes targeted tumor variant analysis via Sanger sequencing, as well as targeted RNA fusion analysis for select pediatric sarcomas. Diagnostic sequencing and screening technologies allow for more refined diagnosis and real-time identification of a subset of patients that benefit from more targeted treatment. Our genetic tests for cancer include:
- Targeted gene fusion based testing for selected malignant sarcomas
- Targeted Sanger sequencing analysis
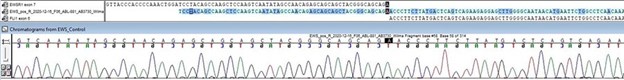
Our laboratory performs cytogenetic chromosome analysis (also known as G-banded analysis or karyotype) to detect single-cell numerical and structural chromosomal abnormalities that may be associated with ultrasound abnormalities, birth defects, developmental delays or disabilities, disorders of sexual differentiation, infertility, pregnancy loss and stillbirth. This test requires fresh specimens (living cells) that can grow and divide in culture to visualize their chromosomes during cell division. We offer chromosome analysis on a variety of specimen types including peripheral blood, skin biopsy, amniotic fluid, periumbilical blood (PUBS), and products of conception.
The image below is a karyogram of a newborn with a balanced translocation between the long arm of chromosome 3 and the long arm of chromosome 12. In addition, there is a terminal deletion of one copy of chromosome 22 consistent with the diagnosis of Phelan-McDermid syndrome.
ISCN : 46,XX,t(3;12)(q27;q23),del(22)(q13.2)
Locus specific probes for well-known microdeletion and microduplication syndromes (e.g. DiGeorge syndrome and Prader–Willi syndrome)
Locus specific probes for familial carrier testing
Locus specific probes for pediatric cancers (e.g. MYCN, MYC and ALK)
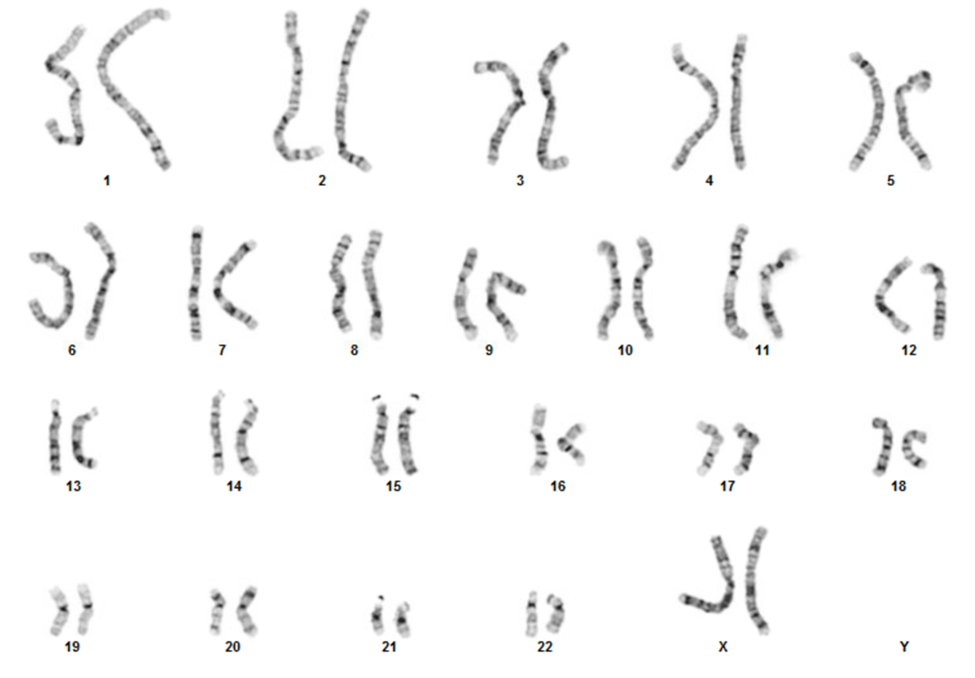
Below, are examples of FISH images. A) MYCN amplification in a snap frozen section of a neuroblastoma tumor; B) Metaphase image demonstrating loss of the DiGeorge syndrome critical region (arrow), also visible in interphase cell showing only one signal for DiGeorge syndrome probe (orange) and two signals for the control probe (green); C) Aneuvysion screen using locus specific probes for chromosome 13 (green) and chromosome 21, (orange). This image depicts fetal cells with an extra copy of chromosome 21.
Chromosomal microarray analysis (CMA) is a genome-wide methodology to identify gains and losses of small or large parts of chromosomes. This method is based on empirically selected single nucleotide polymorphisms (SNPs) spanning the genome with enriched disease-focused coverage. It allows the detection of chromosome gains or losses that are too small to detect with the standard chromosome G-banded testing by microscopy.
This test may also show whether there are two copies from each parent for every chromosome and region. When chromosomes are inherited from one parent and not from the other, it results in large regions of homozygosity (ROH) which can be assessed by single nucleotide polymorphism (SNP) analysis. The presence of ROH is not diagnostic of any disorder, but it can suggest an increased risk for imprinting disorders and recessive genetic disorders.
Chromosomal microarray analysis is frequently ordered for individuals with multiple congenital anomalies and developmental delays. It can also be performed in the prenatal setting for a fetus with ultrasound abnormalities of an unknown cause.
Chromosomal microarray analysis can also be performed on cancer tissue (tumors). Our laboratory tests neuroblastoma and Wilms tumor samples for specific chromosomal gains and losses or segmental aberrations that can inform tumor behavior also known as prognostic indicators.
In addition, our laboratory offers methylation classification of central nervous system (CNS) tumors using CMA technology. This test allows for the detection of methylation signatures that enhances the ability to accurately diagnose these tumors based on their methylation profiles.
Next-generation sequencing (NGS) enables simultaneous analysis of multiple genes in a single comprehensive assay. Our clinical laboratory performs NGS assays for individuals with suspected, but undiagnosed, genetic disease in the setting of Genome Sequencing and Rapid Genome Sequencing for pediatric and adolescent patients. NGS is also used in the setting of cancer to identify gene alterations involved in hematologic malignancies and solid tumors (Paired Somatic Disease-Germline Comparator Exome sequencing, Hematologic Cancer Fusion Analysis, Solid Tumor Fusion Analysis, and Neuroblastoma Panel) for the purpose of diagnosis, prognostication, and therapeutic targeting.
Genome Sequencing
Genome Sequencing (GS) is a method that allows for comprehensive analysis across the entire genome. This test is intended for patients who are suspected to have an underlying genetic disorder, and it is used to identify variants in a patient's DNA that may cause or may be related to their medical condition.
GS can be performed on only the affected patient (proband), or the patient and biological parents. Optimal testing includes sequencing of the patient and both biological parents to determine whether variants are shared with a parent (inherited) or unique to the patient (de novo). Knowledge of parental inheritance is extremely useful for phasing of variants in autosomal recessive conditions, as well as identification of de novo variation. DNA is most frequently obtained for this test from a blood sample. Less frequently, DNA may be obtained from saliva or cheek swabs.
GS can identify variants in a patient’s DNA that may be causal for a patient’s medical condition. Additionally, GS may identify variants that are unrelated to the patient’s medical condition (secondary findings) or reveal information about familial relationships. The American College of Medical Genetics and Genomics (ACMG) has recommendations for reporting secondary findings in a specific subset of genes, which includes genes associated with inherited cancer syndromes and certain types of heart disease. Due to the complex nature of this test, genetic counseling and informed consent are essential parts of the GS testing process.
Oncology Sequencing
Somatic Disease-Germline Comparator Exome sequencing is used in individuals diagnosed with cancer to enable both genetic profiling of the tumor, as well as an assessment of variation associated with cancer predisposition from the germline. This comprehensive assay enables the identification and characterization of genetic variation informing diagnosis, prognostication of disease, and a determination of potential therapeutic targets for cancer treatment. Evaluation for variants associated with cancer predisposition enables appropriate surveillance and management strategies from the clinical care team, along with genetic counseling.
Targeted fusion analysis is performed using either the Hematologic Cancer Fusion Analysis, or Solid Tumor Fusion Analysis, which are RNA-based assays evaluating for the presence of oncogenic gene fusions in hematologic malignancies and solid tumors. A gene fusion is the joining of two or more genes due to a rearrangement. Gene fusions observed in cancers can alter regulatory elements resulting in inappropriate gene expression, or result in aberrant cell signaling, including constitutive activation. Detection of a gene fusion can aid in diagnosis, disease prognostication, and identification of potential therapeutics.
Genetic counselors are health care providers with advanced training in medical genetics and counseling. These providers help guide and support patients who wish to learn about how genetic diseases and conditions can affect them or their families.
Genetic counselors can work in different settings, including a clinical lab like this one. A lab genetic counselor works with other professionals to help ensure best patient care. They do not interact with patients.
Primary roles of genetic counselors at the Institute’s clinical lab at Nationwide Children’s include:
- Serving as a client liaison with providers by answering their questions about appropriate genetic testing
- Reaching out to providers to obtain additional information or clarify test orders
- Ensuring correct test paperwork and samples are provided to the lab
- Coordinating complex testing and ensuring all required needs are met before beginning the test
- Discussing test results with providers and how results may affect other family members of the patient
- Discussing with providers additional testing options for patients if recommended
- Educating community providers about genetic testing offered in the lab
- Helping interpret genetic test results and write lab result reports
- Arranging sending DNA and other samples to outside laboratories for genetic testing
- Working with the clinical lab team to better interpret complex test results to provide the best patient care on an individualized basis
- Helping identify appropriate labs for genetic testing done outside of the lab at Nationwide Children’s
- Helping the clinical lab bring in and implement new test offerings
- Coordinating with the research team to provide appropriate clinical testing for research patients
- Educating students, residents and lab fellows who are training in the lab
Genetic counselors in the laboratory provide support to both our clinical lab team as well as outside providers. While laboratory genetic counselors do not directly interact with patients, they assist in many ways that contribute to direct patient care.

