The American Heart Association (AHA) Institute for Precision Cardiovascular Medicine is dedicated to preserving and prolonging health by harnessing the power of big data to improve outcomes in cardiovascular care. A multidisciplinary research team at Nationwide Children’s Hospital was recently awarded a grant from the AHA, in conjunction with Amazon Web Services, to further this mission through a cloud-based collaboration known as CardioGenomics eXchange commons (CGX), which allows for the analysis and exchange of genomic sequencing data among cardiovascular researchers.
The project team, led by Katie Miller, PhD, postdoctoral scientist in Research Information Solutions and Innovation (RISI) at The Research Institute at Nationwide Children’s, was one of only two awardees out of 80 submissions for the AHA grant. Simon Lin, MD, MBA, chief research information officer at The Research Institute, is the principal investigator for the project, and Vidu Garg, MD, director of the Center for Cardiovascular Research at The Research Institute and physician in The Heart Center at Nationwide Children’s, is co-principal investigator. Additional members of the research team include Kim McBride, MD, MS, principal investigator in the Center for Cardiovascular Research at The Research Institute and division chief of Molecular and Human Genetics; Soheil Moosavinasab, MS, data scientist in RISI; Matt Bailey, full stack web developer on the RISI R&D team; and Ashley Kubatko, MS, PMP project manager for The Research Institute.
“Specifically, our research is aimed at identifying gene mutations responsible for human heart disease,” explains Dr. Miller. “Research efforts have revealed rare, single base changes in DNA that can explain why some individuals have heart disease. We have developed a web application called Genome Dashboard that allows researchers or clinicians to upload genome sequencing data and interactively explore the data to identify gene variants, if any, that may be causal for a patient’s disease.”
The research team integrated various informatics resources from existing public databases, such as Online Mendelian Inheritance in Man (OMIM) and ClinVar, a freely accessible, public archive of reports of the relationships among human variations and phenotypes. These resources provide information about the clinical interpretation of specific genomic findings in humans and animal models, which will allow users to upload, explore, and interpret sequencing data against known disease gene associations. The team will then merge Genome Dashboard into the cloud-based American Heart Association Precision Medicine Platform using Amazon Web Services to create a genomics tool called CardioGenomics eXchange commons, or CGX.
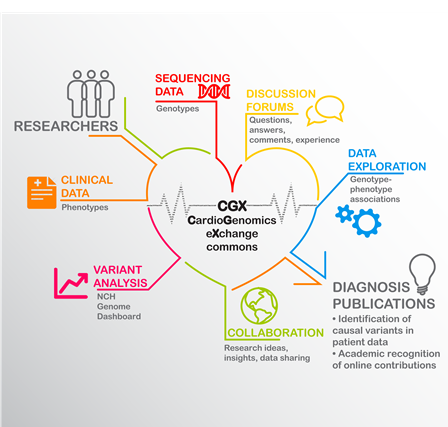
“The CGX will allow users to upload, analyze and share genomic sequencing data and communicate research and clinical findings related to heart disease and precision medicine,” says Dr. Lin. “It will also provide a valuable, necessary resource for the cardiovascular research community to pursue research avenues that might otherwise be dismissed or impossible, due to limitations in aggregation, access, or use of large genomic datasets and clinical data.”
The AHA competitive grants and fellowships foster cross-disciplinary learning, research and collaboration by bringing together scientists with unique backgrounds in data science and computer engineering. As part of the AHA Data Grant Portfolio, Amazon Web Services (AWS) will provide grant recipients, like Lin, credits for computational analysis and data storage on the AHA Precision Medicine Platform powered by Amazon Web Services.
“The promise of precision cardiovascular medicine and care can be realized when research and technology come together to deliver new insights,” said Jennifer Hall, PhD, Chief of the AHA Institute for Precision Cardiovascular Medicine. “The AHA and AWS collaboration will unite the global research community to accelerate discovery in cardiovascular health and usher in a new era of tailored prevention and treatment that will help patients and lessen the global burden of cardiovascular disease.”
According to the team, harnessing the collective intelligence of many researchers and clinicians is not typically leveraged like this in the analysis of genomic data. Their hope is that identifying gene mutations in cardiac patients could ultimately help tailor disease treatments for individuals or groups of patients, an approach known as “precision medicine.” Additionally, the team anticipates that this project will inform and inspire the design of future large-scale tools for data sharing among research communities.
The team looks to synch CGX with the AHA Precision Medicine Platform which would be a boon for researchers and clinicians. The AHA Precision Medicine Platform, powered by Amazon Web Services, is a cloud-based data marketplace where scientists from around the world can access, store, share and analyze research data. For more information on the Precision Medicine Platform, visit http://precision.heart.org.
